MANTISSA
Massive Acceleration of New Techniques In Science with Scalable Algorithms
Motivation
Scalable Statistics and Machine Learning Algorithms are essential for extracting insights from Big Data. Our interdisciplinary team is trying to address a number of challenging analysis problems from a number of science domains at Lawrence Berkeley National Lab (Berkeley Lab). We are developing novel algorithms, and using state-of-the-art methods from High-Performance Computing, Scientific Data Management, and Parallel I/O for running software at scale on NERSC resources.
Problem Domains
| Science Domain | Data Source | Data Characteristics | Dataset Size | Analysis Requirements |
|---|---|---|---|---|
| DESI | Multiple Telescopes | Multi-band, Noisy, Artifacts | O(100)TB | Data fusion, Inference |
| Daya Bay | Detectors | Spatio-Temporal, Noisy, Artifacts | O(100)TB | Pattern/Anomaly Detection |
| Life Sciences | Mass Spectroscopy Imaging | Multi-Modal, Noisy, Artifacts | O(1)TB | Dim. Reduction, Clustering |
| Genomics | Sequencers | Missing Data, Acquisition errors | O(1-10)TB | Clustering, Pattern Detection |
| Climate | Satellites, Simulations | Multi-Variate, Spatio-Temporal | O(100)TB | Pattern Detection |
DESI (Astronomy)

We are addressing the problem of extracting a unified object catalog of objects in the sky from multiple telescope datasets (Sloan Digital Sky Survey, Palomar Transient Factory, Wide-Field Infrared Survey Explorer). The catalog (with associated uncertainties) will be used for target selection phase for the DESI project. We are formulating the problem as a graphical model; the major challenge is in performing efficient inference on the trillion-pixel dataset. We will explore stochastic variational inference and distributed Markov Chain Monte Carlo approaches for parameter estimation.
Daya Bay

Daya Bay is a reactor neutrino experiment located in China, with a host of international collaborators. A range of anti-neutrino detectors capture signatures corresponding to neutron decay, supernovae explosions, cosmogenic activity and background radiation. The overall dataset is more than 500TB. Current approaches for analysis of the Daya-Bay time-series consists of a manual exploration of short time windows. We are interested in applying unsupervised machine learning technology for automatically inferring high level categories of event types.
Life Sciences (Mass Spectroscopy Imaging)
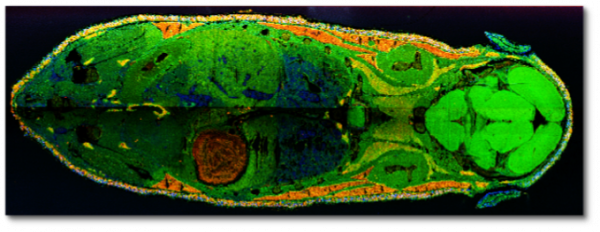 Mass-spec imaging datasets provide unprecedented fidelity in measuring spatial distribution and abundances of molecules at locations within a sample. MSI has widespread applications in life sciences, bio-engineering, medical diagnosis and scientific experiments. Unfortunately, the lack of scalable data analysis capabilities is hindering progress in this field. We are exploring randomized linear algebra techniques for processing raw TB-sized MSI data volumes consisting of billions of voxels. We are exploring CUR decomposition and Nystrom approximations of spatial kernels; with the goal of producing scientifically interpretable basis vectors and meaningful clusters.
Mass-spec imaging datasets provide unprecedented fidelity in measuring spatial distribution and abundances of molecules at locations within a sample. MSI has widespread applications in life sciences, bio-engineering, medical diagnosis and scientific experiments. Unfortunately, the lack of scalable data analysis capabilities is hindering progress in this field. We are exploring randomized linear algebra techniques for processing raw TB-sized MSI data volumes consisting of billions of voxels. We are exploring CUR decomposition and Nystrom approximations of spatial kernels; with the goal of producing scientifically interpretable basis vectors and meaningful clusters.
Genomics
 Graph-theoretical methods are prevalent in genomics and forms the basis of genetic mapping, gene-expression analysis, and genome assembly. Constructing graphs from the input data involve computing pairwise similarities and runs in at least O(n2k) time for n points with k features. The graph construction can hence be costlier than subsequent analysis. We will explore methods to compute important graph kernels such as minimum spanning forests directly on the input data itself without computing all pairwise similarities. We are exploring metric data structures as well as techniques that exploit the underlying low dimensional geometry.
Graph-theoretical methods are prevalent in genomics and forms the basis of genetic mapping, gene-expression analysis, and genome assembly. Constructing graphs from the input data involve computing pairwise similarities and runs in at least O(n2k) time for n points with k features. The graph construction can hence be costlier than subsequent analysis. We will explore methods to compute important graph kernels such as minimum spanning forests directly on the input data itself without computing all pairwise similarities. We are exploring metric data structures as well as techniques that exploit the underlying low dimensional geometry.
Climate
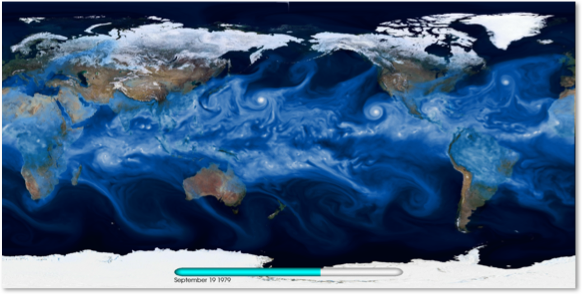 We are interested in mining multi-variate, spatio-temporal climate datasets for extreme weather patterns (Hurricanes, Extra-Tropical Cyclones, Atmospheric Rivers, Heatwaves, Blocking events, etc). High-resolution climate datasets produce O(100) TB output per run; current CMIP-5 archives consist of 5PB datasets, posing major scalability challenges for analysis tools. We will explore DBN methodologies and evaluate their performance for supervised learning tasks on climate datasets. We will also extend these methods for operating on multi-variate datasets.
We are interested in mining multi-variate, spatio-temporal climate datasets for extreme weather patterns (Hurricanes, Extra-Tropical Cyclones, Atmospheric Rivers, Heatwaves, Blocking events, etc). High-resolution climate datasets produce O(100) TB output per run; current CMIP-5 archives consist of 5PB datasets, posing major scalability challenges for analysis tools. We will explore DBN methodologies and evaluate their performance for supervised learning tasks on climate datasets. We will also extend these methods for operating on multi-variate datasets.
Project Team
Researchers
- Aydin Buluç, Ben Bowen (Berkeley Lab)
- Michael Mahoney, Jiyang Yang (UC-Berkeley/ICSI/Stanford)
- Jon McAuliffe, Jeff Regier, Fritz Sommer, Urs Koester (UC-Berkeley Stats/Redwood)
- Ryan Adams, Brenton Patridge, Oren Rippel, Albert Wu, Elaine Angelos (Harvard/MIT)
Berkeley Lab Collaborators
- David Schlegel, Dan Dwyer (Physics)
- Peter Nugent, Craig Tull, Michael Wehner, Oliver Ruebel, Lenny Oliker (Applied Mathematics & Computational Research)
- Wahid Bhimji, Evan Racah (NERSC)
- Trent Northern, Katherine Louie (Life Sciences)
- Dan Roshkar, Jarrod Chapman (Joint Genome Institute)
External Collaborators
- Joey Gonzalez, Rashmi K V (UCB AMPLab)
- Matt Hoffman (Adobe Research)
- Dustin Lang (CMU)
- Mohit Singh, Nadathur Satish, Narayanan Sundaram, Mostafa Patwari (Intel Research)
- Amir Khosorowshahi







