A Game Changer: Metagenomic Clustering Powered by HPC
New Berkeley Lab algorithm allows biologists to harness the capabilities of massively parallel supercomputers to make sense of a genomic ‘data deluge’
March 12, 2018
By Linda Vu
Contact: cscomms@lbl.gov
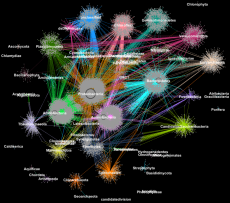
Proteins from metagenomes clustered into families according to their taxonomic classification. (Credit: Georgios Pavlopoulos and Nikos Kyrpides, JGI/Berkeley Lab)
Download HipMCL.
Did you know that the tools used for analyzing relationships between social network users or ranking web pages can also be extremely valuable for making sense of big science data? On a social network like Facebook, each user (person or organization) is represented as a node and the connections (relationships and interactions) between them are called edges. By analyzing these connections, researchers can learn a lot about each user—interests, hobbies, shopping habits, friends, etc.
In biology, similar graph-clustering algorithms can be used to understand the proteins that perform most of life’s functions. It is estimated that the human body alone contains about 100,000 different protein types, and almost all biological tasks—from digestion to immunity—occur when these microorganisms interact with each other. A better understanding of these networks could help researchers determine the effectiveness of a drug or identify potential treatments for a variety of diseases.
Today, advanced high-throughput technologies allow researchers to capture hundreds of millions of proteins, genes and other cellular components at once and in a range of environmental conditions. Clustering algorithms are then applied to these datasets to identify patterns and relationships that may point to structural and functional similarities. Though these techniques have been widely used for more than a decade, they cannot keep up with the torrent of biological data being generated by next-generation sequencers and microarrays. In fact, very few existing algorithms can cluster a biological network containing millions of nodes (proteins) and edges (connections).
That’s why a team of researchers from the Department of Energy’s (DOE’s) Lawrence Berkeley National Laboratory (Berkeley Lab) and Joint Genome Institute (JGI) took one of the most popular clustering approaches in modern biology—the Markov Clustering (MCL) algorithm—and modified it to run quickly, efficiently and at scale on distributed-memory supercomputers. In a test case, their high-performance algorithm—called HipMCL—achieved a previously impossible feat: clustering a large biological network containing about 70 million nodes and 68 billion edges in a couple of hours, using approximately 140,000 processor cores on the National Energy Research Scientific Computing Center’s (NERSC) Cori supercomputer. A paper describing this work was recently published in the journal Nucleic Acids Research.
“The real benefit of HipMCL is its ability to cluster massive biological networks that were impossible to cluster with the existing MCL software, thus allowing us to identify and characterize the novel functional space present in the microbial communities,” says Nikos Kyrpides, who heads JGI’s Microbiome Data Science efforts and the Prokaryote Super Program and is co-author on the paper. “Moreover we can do that without sacrificing any of the sensitivity or accuracy of the original method, which is always the biggest challenge in these sort of scaling efforts.”
“As our data grows, it is becoming even more imperative that we move our tools into high performance computing environments, ” he adds. “If you were to ask me how big is the protein space? The truth is, we don’t really know because until now we didn’t have the computational tools to effectively cluster all of our genomic data and probe the functional dark matter.”
In addition to advances in data collection technology, researchers are increasingly opting to share their data in community databases like the Integrated Microbial Genomes & Microbiomes (IMG/M) system, which was developed through a decades-old collaboration between scientists at JGI and Berkeley Lab’s Computational Research Division (CRD). But by allowing users to do comparative analysis and explore the functional capabilities of microbial communities based on their metagenomic sequence, community tools like IMG/M are also contributing to the data explosion in technology.
How Random Walks Lead to Computing Bottlenecks

Cori Supercomputer at the National Energy Research Scientific Computing Center (NERSC). (Photo by Roy Kaltschmidt, Berkeley Lab)
To get a grip on this torrent of data, researchers rely on cluster analysis, or clustering. This is essentially the task of grouping objects so that items in the same group (cluster) are more similar than those in other clusters. For more than a decade, computational biologists have favored MCL for clustering proteins by similarities and interactions.
“One of the reasons that MCL has been popular among computational biologists is that it is relatively parameter free; users don’t have to set a ton of parameters to get accurate results and it is remarkably stable to small alterations in the data. This is important because you might have to redefine a similarity between data points or you might have to correct for a slight measurement error in your data. In these cases, you don’t want your modifications to change the analysis from 10 clusters to 1,000 clusters,” says Aydin Buluç, a CRD scientist and one of the paper’s co-authors.
But, he adds, the computational biology community is encountering a computing bottleneck because the tool mostly runs on a single computer node, is computationally expensive to execute and has a big memory footprint—all of which limit the amount of data this algorithm can cluster.
One of the most computationally and memory intensive steps in this analysis is a process called random walk. This technique quantifies the strength of a connection between nodes, which is useful for classifying and predicting links in a network. In the case of an Internet search, this may help you find a cheap hotel room in San Francisco for spring break and even tell you the best time to book it. In biology, such a tool could help you identify proteins that are helping your body fight a flu virus.
Given an arbitrary graph or network, it is difficult to know the most efficient way to visit all of the nodes and links. A random walk gets a sense of the footprint by exploring the entire graph randomly; it starts at a node and moves arbitrarily along an edge to a neighboring node. This process keeps going until all of the nodes on the graph network have been reached. Because there are many different ways of traveling between nodes in a network, this step repeats numerous times. Algorithms like MCL will continue running this random walk process until there is no longer a significant difference between the iterations.
In any given network, you might have a node that is connected to hundreds of nodes and another node with only one connection. The random walks will capture the highly connected nodes because a different path will be detected each time the process is run. With this information, the algorithm can predict with a level of certainty how a node on the network is connected to another. In between each random walk run, the algorithm marks its prediction for each node on the graph in a column of a Markov matrix—kind of like a ledger—and final clusters are revealed at the end. It sounds simple enough, but for protein networks with millions of nodes and billions of edges, this can become an extremely computationally and memory intensive problem. With HipMCL, Berkeley Lab computer scientists used cutting-edge mathematical tools to overcome these limitations.
“We have notably kept the MCL backbone intact, making HipMCL a massively parallel implementation of the original MCL algorithm,” says Ariful Azad, a computer scientist in CRD and lead author of the paper.
Although there have been previous attempts to parallelize the MCL algorithm to run on a single GPU, the tool could still only cluster relatively small networks because of memory limitations on a GPU, Azad notes.
“With HipMCL we essentially rework the MCL algorithms to run efficiently, in parallel on thousands of processors, and set it up to take advantage of the aggregate memory available in all compute nodes,” he adds. “The unprecedented scalability of HipMCL comes from its use of state-of-the-art algorithms for sparse matrix manipulation.”
According to Buluç, performing a random walk simultaneously from many nodes of the graph is best computed using sparse-matrix matrix multiplication, which is one of the most basic operations in the recently released GraphBLAS standard. Buluç and Azad developed some of the most scalable parallel algorithms for GraphBLAS’s sparse-matrix matrix multiplication and modified one of their state-of-the-art algorithms for HipMCL.
“The crux here was to strike the right balance between parallelism and memory consumption. HipMCL dynamically extracts as much parallelism as possible given the available memory allocated to it,” says Buluç.
HipMCL: Clustering at Scale
In addition to the mathematical innovations, another advantage of HipMCL is its ability to run seamlessly on any system—including laptops, workstations and large supercomputers. The researchers achieved this by developing their tools in C++ and using standard MPI and OpenMP libraries.
“We extensively tested HipMCL on Intel Haswell, Ivy Bridge and Knights Landing processors at NERSC, using a up to 2,000 nodes and half a million threads on all processors, and in all of these runs HipMCL successfully clustered networks comprising thousands to billions of edges,” says Buluç. “We see that there is no barrier in the number of processors that it can use to run and find that it can cluster networks 1,000 times faster than the original MCL algorithm.”
“HipMCL is going to be really transformational for computational biology of big data, just as the IMG and IMG/M systems have been for microbiome genomics,” says Kyrpides. “This accomplishment is a testament to the benefits of interdisciplinary collaboration at Berkeley Lab. As biologists we understand the science, but it’s been so invaluable to be able to collaborate with computer scientists that can help us tackle our limitations and propel us forward.”
Their next step is to continue to rework HipMCL and other computational biology tools for future exascale systems, which will be able to compute quintillion calculations per second. This will be essential as genomics data continues to grow at a mindboggling rate—doubling about every five to six months. This will be done as part of DOE Exascale Computing Project’s Exagraph co-design center.
Development of HipMCL was primarily supported by the U.S. Department of Energy’s Office of Science via the Exascale Solutions for Microbiome Analysis (ExaBiome) project, which is developing exascale algorithms and software to address current limitations in metagenomics research. The development of the fundamental ideas behind this research was also supported by the Office of Advanced Scientific Computing Research’s Applied Math Early Career program. NERSC is a DOE Office of Science User Facility.
HipMCL is based on MPI and OpenMP and is freely available under a modified BSD license. Installation instructions and large datasets can be found in the relevant Wiki tab.
About NERSC and Berkeley Lab
The National Energy Research Scientific Computing Center (NERSC) is a U.S. Department of Energy Office of Science User Facility that serves as the primary high performance computing center for scientific research sponsored by the Office of Science. Located at Lawrence Berkeley National Laboratory, NERSC serves almost 10,000 scientists at national laboratories and universities researching a wide range of problems in climate, fusion energy, materials science, physics, chemistry, computational biology, and other disciplines. Berkeley Lab is a DOE national laboratory located in Berkeley, California. It conducts unclassified scientific research and is managed by the University of California for the U.S. Department of Energy. »Learn more about computing sciences at Berkeley Lab.







