Turning Grass into Gas for Less
Simulations Unlock Secrets of Cellulose Deconstruction
September 30, 2011
By Margie Wylie
Contact: cscomms@lbl.gov

If researchers can unlock the sugars stored in plant cellulose, then crops like this switchgrass could be turned into biofuels, rather than using corn or other food crops.
Pull up to the pump these days, and chances are your gas will be laced with ethanol, a biofuel made from corn. Corn-ethanol is relatively easy to make, but with growing populations and shrinking farmland, there will never be enough of the starchy food crop to both feed and fuel the world.
That’s why researchers are working on “grassoline,” liquid biofuels made from hardy, high-yielding, non-food crops like switchgrass. But what makes these crops indigestible to humans also makes them challenging raw materials for biofuel production. The sugars needed to make biofuels are locked up tight in cellulose, and researchers have yet to figure out an economical, scalable way to break them loose.
Recent computer simulations carried out at the National Energy Research Scientific Computing Center (NERSC) could help scientists do just that. The simulations show how two different solvents help unzip cellulose’s structure, peeling off long strings of sugars. That information should help researchers engineer molecules that do the job better, cheaper, and on a larger scale, says Jhih-Wei Chu, a chemical and biomolecular engineering professor at the University of California Berkeley who ran the simulations at NERSC.
Computer simulations offer scientists an unparalleled view into processes that are practically unobservable. “If you try to measure cellulose degradation experimentally, it’s just too messy: The signals overlap and it’s impossible to isolate the individual forces and positions of molecules,” he explains. “With simulations, we can calculate atom-by-atom, bond-by-bond exactly what is happening in a complex molecular system.” Simulations also offer researchers the luxury of studying few billionths-of-a-second processes at human time scales, said Chu, who is also a principal investigator with the Energy Biosciences Institute (EBI). EBI is a collaboration of the university, the U.S. Department of Energy’s Lawrence Berkeley National Lab (which operates NERSC), and the energy company BP.
Cracking Cellulose
Cellulose is a polymer of the sugar glucose, the most important energy source for living things. Plants make glucose from sunlight, water, and carbon dioxide through the process of photosynthesis. They link glucose molecules together with oxygen, forming long chains. Then, the chains are aligned into sheets that are bound together with hydrogen bonds to form microfibrils of cellulose.
Humans now want to take that same building material and convert it back into energy. The problem is that cellulose is very tough stuff. “Plants have evolved to synthesize cellulose to resist breakdown,” Chu said.
Still, cellulose isn’t impervious to degradation. Toss an old cotton rag (about 90 percent cellulose) into your garden, and it will eventually degrade like other plant material, but it can take months or years. Scientists and engineers who work with biofuels use a variety of methods to help speed the process along, including steam, pressure, enzymes, and solvents.
Chu simulated two aspects of cellulose decomposition: solvent effects and enzyme actions.
Solvent Effects
In his first simulation study, Chu explored how liquid salts (ionic liquids, in scientific parlance) dissolve crystalline cellulose. Sodium chloride, common table salt, has a very compact crystalline structure so doesn’t turn liquid until it reaches 800 degrees Centigrade. To get a salt that’s liquid at much lower temperatures, the sodium cation (the positively charged ion) is replaced with a bulkier organic cation.
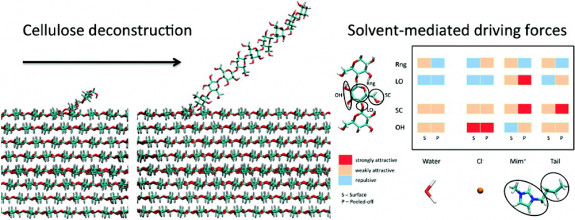
On the left, a strand of cellulose is peeled off from the crystalline structure. On the right, the table documents the attractive forces between various components of cellulose and the ionic liquid. Select to enlarge. (Credit: Jhih-Wei Chu, UC Berkeley)
For his simulations, Chu used 1-butyl-3-methylimidazolium chloride (BmimCl). “We know this ionic liquid will dissolve cellulose, but we are really not sure why,” says Chu. “Along with my co-authors, I wanted to understand how the molecular forces provided by ionic liquids help deconstruct crystalline cellulose. That knowledge could allow us to design better solvents and engineering processes.”
In the lab, there are many ionic liquids to choose from. That’s the good news. The bad news is that there are so many ionic liquids to choose from that it is impractical for researchers to experiment with all possible candidates in a trial-and-error fashion. “If there are N cations and N anions to choose from, then there can be up to N2 possible ionic liquids. That’s a lot to sort through,” said Chu. “To rationally design solvents for cellulose, you have to know what properties you are after and how the ionic liquid interacts with cellulose. Therefore, a powerful ‘microscope’ with the ability to resolve molecular structures and also sense their interaction forces is needed. That is where computer simulation comes in.”
His simulation model, which contains approximately 100,000 clusters of atoms, required around one million CPU hours split between the Hopper system (a Cray XE6 supercomputer) and Carver (an IBM iDataplex cluster) to produce a few tens of billionths of a second of action. “The range of systems at NERSC is great for our work,” says Chu. “We run these enormous calculations on Hopper, but for some theory, the amount of memory per node is more important than the number of cores you can run at once and that’s where Carver comes in.”
Chu’s model breaks down the cellulose polymer into four different sections – the hexagonal ring (Rng), the linker oxygen (LO), the side chain (SC), and the hydroxyl groups (OH). In turn, the interactions of these groups were tested against water, the anion (Cl-), the charged portion of the cation (Mim+), and the neutral tail portion of the cation (Tail). The table (above) summarizes the interactions between all these groups for a strand of cellulose before and after the solvent peels it off.
If the various components of cellulose are more attracted to each other than to the solvent, cellulose won’t dissolve. This is the case with water, a poor solvent for cellulose.
However, in BmimCl, the components of cellulose are more attracted to the components of the ionic liquid than they are to each other, and the cellulose dissolves.
Chu and collaborators found that this solvent is very versatile, attacking a variety of bonds in the crystalline structure. “The Cl anions have strong attraction to hydroxyl groups and weak attraction to side chains and sugar rings, while the cations have strong attractions to side chains and linker oxygens,” said Chu. “This versatility can potentially be employed as the basis for molecular design and engineering.”
In a second simulation study, Chu and colleagues examined the workings of a small portion of a much more complex system, a cellulase enzyme.
Enigmatic Enzymes
Evolution has endowed some organisms with cellulases, enzymes that can break down cellulose. Cow and termites digest cellulose, although they rely on cellulases of symbiotic bacteria living in the acidic environment of their digestive tracts. Humans can’t digest the cellulose in our food. We call it fiber.
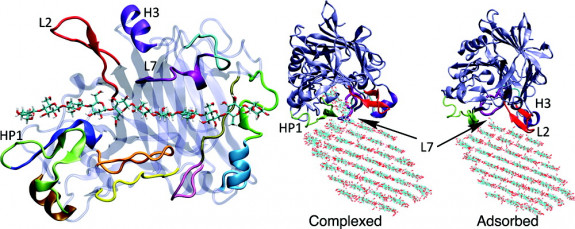
A strand of cellulose passes through the active site of the Trichoderma reesei cellulase enzyme. One portion of the enzyme, marked L7, acts like an ice cream scoop, physically peeling up a fibril and lifting it into the active site for digestion. Select to enlarge. (Credit: Jhih-Wei Chu, UC Berkeley)
One of the best sources of cellulases is the fungus Trichoderma reesei. During WWII in the South Pacific, the U.S. Army discovered that its canvas tents were dissolving in the heat and humidity. Army researchers isolated the fungus feeding on their tents, and it has been an important source of cellulases ever since.
Like ionic liquids, researchers have been using cellulases to degrade cellulose without entirely understanding how they work.
In a recent simulation, Chu and collaborators found that a portion of a T. reesei cellulase acts something like an ice cream scoop, physically peeling up microfibrils of cellulose and lifting them into the maw of the enzyme for digestion. The simulation hints at ways industrial enzymes already in use might be improved, said Chu.
“Computer simulation of molecular dynamics and interactions really is like a huge crowd shot in a movie. As the director, there is so much detail at your disposal that you can lose sight of the plot. Knowing where to zoom in requires some judgment and intuition,” said Chu. “I think we were able to distill the essential plot elements that we hope biofuel researchers will be able to develop further.”
“Dissecting Force Interactions in Cellulose Deconstruction Reveals the Required Solvent Versatility for Overcoming Biomass Recalcitrance” appeared in the Journal of the American Chemical Society on July 28, 2011.
“Protein Allostery at the Solid-Liquid Interface: Endoglucanase Attachment to Cellulose Affects Glucan Clenching in the Binding Cleft” appeared in the Journal of the American Chemical Society on August 30, 2011.
About NERSC and Berkeley Lab
The National Energy Research Scientific Computing Center (NERSC) is a U.S. Department of Energy Office of Science User Facility that serves as the primary high performance computing center for scientific research sponsored by the Office of Science. Located at Lawrence Berkeley National Laboratory, NERSC serves almost 10,000 scientists at national laboratories and universities researching a wide range of problems in climate, fusion energy, materials science, physics, chemistry, computational biology, and other disciplines. Berkeley Lab is a DOE national laboratory located in Berkeley, California. It conducts unclassified scientific research and is managed by the University of California for the U.S. Department of Energy. »Learn more about computing sciences at Berkeley Lab.







