New Insights Into the Human Gut Microbiome
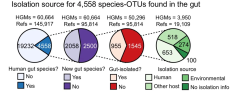
Of the 23,790 species-level OTUs identified from MAGs and reference genomes, 4,558 were classified as being from the human gut on the basis of (1) having a MAG from the HGM dataset, (2) being detected in a human gut metagenome via read-mapping with IGGsearch or (3) containing a reference genome with metadata that indicate isolation from a human stool sample. Of the 4,558 gut OTUs, 2,058 are represented exclusively by MAGs from the current study and are therefore newly identified. Of the remaining 2,500 represented by reference genomes, only 955 contained a gut-isolated reference genome. The remaining 1,545 OTUs either lack isolation metadata or contain metadata that indicate other isolation sources, including human, non-human and environmental. For example, several gut species from non-host-associated environments were isolated from human food products, including milk, cheese, meat and fermented foods.
Science Achievement
Scientists at the DOE Joint Genome Institute computationally reconstructed 60,664 microbial genomes from 3,810 human gut metagenomes from a diverse set of human subjects. These genomes represent 2,058 previously unknown species, thereby bringing the number of known human gut species to 4,558 and increasing the diversity of sequenced gut bacteria by 50 percent.
Significance and Impact
The gut microbiome plays a myriad of important roles in human health and disease. Microbial reference genomes are essential resources for understanding the functional role of specific organisms and for quantifying their abundance from metagenomes. However, an estimated 40-50% of human gut species lack a reference genome, largely because these organisms have not been isolated under laboratory conditions.
This dataset is expected to be used to guide future culturing efforts in the human gut microbiome. The team identified numerous large, uncultivated human gut lineages that could be prioritized for cultivation. Further, they identified genes and pathways that are commonly lost from uncultivated bacteria, which may point towards new growth factors. The collection of 60,664 genomes and the new microbiome profiling tool, IGGsearch, will be useful resources for the human microbiome community and should promote further discoveries in this important microbial community.
Research Details
The researchers developed a computational tool to identify the abundance of all 4,558 human gut species. Using this tool, they compared the microbiome between healthy and diseased individuals and identified 2,283 associations for 10 different diseases. Nearly 40% of species-disease associations correspond to the 2,058 new species, indicating that the current study has provided a more complete picture of how the microbiome is involved in various human diseases.
To address this question of why so many human gut species are unknown, the researchers compared reconstructed genomes between species that have been cultivated to those that have not. They found that uncultivated species have genomes that are on average 19% smaller and are missing numerous genes for biosynthesis of fatty acids, amino acids, and vitamins. These gene losses may indicate important growth factors that are not included in currently used growth media.
The project extensively used large scale computing on the Denovo and Cori NERSC systems. It required nearly 10 million files, 40 TB of disk space and more than 1 million compute hours.
Related Links
Stephen Nayfach, Zhou Jason Shi, Rekha Seshadri, Katherine S. Pollard & Nikos C. Kyrpides, "New insights from uncultivated genomes of the global human gut microbiome"; Nature volume 568, pages 505-510 (2019) , 10.1038/s41586-019-1058-x, NERSC repository: m342
About NERSC and Berkeley Lab
The National Energy Research Scientific Computing Center (NERSC) is a U.S. Department of Energy Office of Science User Facility that serves as the primary high performance computing center for scientific research sponsored by the Office of Science. Located at Lawrence Berkeley National Laboratory, NERSC serves almost 10,000 scientists at national laboratories and universities researching a wide range of problems in climate, fusion energy, materials science, physics, chemistry, computational biology, and other disciplines. Berkeley Lab is a DOE national laboratory located in Berkeley, California. It conducts unclassified scientific research and is managed by the University of California for the U.S. Department of Energy. »Learn more about computing sciences at Berkeley Lab.







